1.7.2 Profiling
We have experimented with some very simple code profiling by introducing the time() function into our code in previous examples using it to record the start and end times of our code and check the overall performance.
While that gives us a high level view of the performance of our code, we do not know where specific bottlenecks might exist within that code. For example what is the slowest part of our algorithm, is it the file reading or writing or the calculations in between? When we know where these bottlenecks are we can investigate ways of removing them or at least using faster techniques to speed up our code.
In this section, we will focus on basic profiling that looks at how long each function call takes – not each individual line of code. This basic type of code profiling is built into Spyder (and most IDEs). However, we also provide some complementary materials that explain how to visualize profiling results and on how to profile each line of code individually but these parts will be entirely optional because they are quite a bit more complex and require the installation of additional software and packages. It is possible that you will run into some technical/installation issues, so our recommendation is that you only come back and try to run the described steps in these optional subsections yourself if you are done with lesson and homework assignment and still have time left.
1.7.2.1 Basic code profiling
We will again use the Spyder IDE and our basic raster code from earlier in the lesson. Spyder has a Profile pane as well which is accessible from the View -> Panes menu. You may need to manually load your code into Profiler using the folder icon. The spyder help for the Profiler is here if you'd like to read it (Profiler — Spyder 5 documentation (spyder-ide.org) [1]) but we explain the important parts below.
Once you load your code, Profiler automatically starts profiling it and displays the message “ Profiling, please wait...“ at the top of the Profiler window. You will need to be a little patient as Spyder runs through all of your code to perform the timing (or alternatively in these raster examples reduce the sample size). You probably remember that we recommended to run multiprocessing code from the command line rather than inside Spyder. However, using the built-in profiler for loading and running multiprocessing code works as long as the worker function is defined in its own module as we did for the vector clipping example to be able to use it as a script tool. If this is not the case, you will receive an error that certain elements in the code cannot be “pickled“ which you might remember from our multiprocessing discussion means those objects cannot be converted to a basic type. We didn't split the multiprocessing version of the raster example into two separate modules, so here we will only look at the non-multiprocessing version and profile that. We won't have this issue when we using other profilers in the following optional sections and in the homework assignment you will work with the vector clipping multiprocessing example which has been set up in a way that allows for profiling with the Spyder profiler.
Once the Profiler has completed you will see a set of results like the ones below.
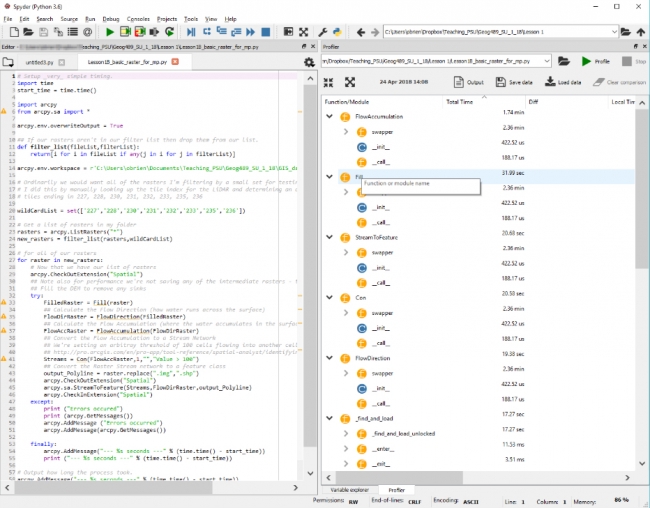
Looking over those results you will see a list of functions together with the times each has taken. The important column to examine is the Local Time column which shows you how long each function has taken to execute in us (microseconds), ms (milliseconds), seconds, minutes etc. The Total Time column is showing you the cumulative time for each of those processes that was run (e.g. if your code was running in a function). You can order any of the columns but if you arrange the Total Time column in ascending order this will give you a logical starting point as the times will be arranged from the shortest to longest running. There is no way to order the results to see the order in which your code ran. So you will see (depending on your code arrangement) overwriteOutput followed by the time function, then filterList, etc.
The next column to look at is the Calls column which has the count of how many times each of those functions was launched. So a high value in Local Time might be indicative of either a large number of calls to a very fast function or a small number of calls to a very slow function.
In my timings, there aren’t any obvious places to look for performance improvements although the code could be fractionally faster (but less of a good team player) if I didn’t check the extension back in and my .replace() method and print add a small amount of time to the execution.
What we are doing with this type of profiling is examining the sum of the functions and methods which were called during the code execution and how long they took, in total, for the number of times that they were called. It is possible to identify inefficiencies with this sort of profiling, particularly in combination with debugging. Am I calling a slow function too often? Can I find a faster function to do the same job (for example some mathematical functions are significantly faster than others and achieve the same result and often there exist approximations that give almost the exact result but are much faster to compute)?
It is worth pointing out here that the results from Spyder’s Profiler are actually the output from the cProfile [2] package of the Python standard library which is essentially wrapped around our script to calculate the statistics we are seeing above. You could import this package into your own code and use it there directly but we will focus on using its functionality from the IDE which is usually more convenient and the results are presented in a more readily understood format.
You might be thinking that these results aren’t really that readily understood and it would be easier if there were a graphical visualization of the timings. Luckily there is and if you want to learn more about it, the following optional sections on code profiling with visualiuation are a good starting point. In addition, we continue another optional section that explains how you can do more detailed profiling looking at each individual line rather than complete functions. However, we recommend that you skip or only skim through these optional sections on your first pass through the lesson materials and come back when you have the time.
1.7.2.2 Optional complementary materials: Code profiling with visualizations
As we said at the beginning of this section, this section is provided for interest only. Please feel free to skip over it and you can loop back to it at the end of the lesson if you have free time or after the end of the class. Be warned: installing the required extensions and using them is a little complicated - but this is an advanced class, so don't let this deter you.
We are going to need to download some software called QCacheGrind which reads a tree type file (like a family tree). Unfortunately, QcacheGrind doesn’t natively support the profile files we are going to be creating so we will also need a converter (pyprof2calltree [3]), written in Python. Our workflow is going to be :
- Download & install QCacheGrind (we only need to do this once)
- Use pip (a Python package manager) to install our converter (we only need to do this once, too)
- Run our function and line profiling and save the output to files
- Convert those output profile files using our converter
- Open the converted files in QCacheGrind
- ...
- Conquer Python (okay maybe not, but at least have a better understanding of our code’s performance)
Installing QCacheGrind
Download QCacheGrind [4] and unzip it to a folder. QcacheGrind can be run by double-clicking on the qcachegrind executable in the folder you’ve just unzipped it to. Don’t do that just yet though, we’ll come back to it once we’ve done the other steps in our workflow and when we have some profile files to visualize.
Installing the Converter - pyprof2calltree
Now we’re going to install our converter using the Python Preferred Installer Program, pip. If you would like to learn more about pip, the Python 3 Help (Key Terms) [5] has a full explanation. You will also learn more about Python packet managers in the next lesson. Pip is included by default with the Python installation you have but we have to access it from the Python Command Prompt.
As we mentioned in Section 1.6.5.2, there should be a shortcut within the ArcGIS program group on the start menu called "Python Command Prompt" on your PC that opens a command window running within the conda environment indicating that this is Python 3 (py3). You actually may have several shortcuts with rather similar sounding names, e.g. if you have both ArcGIS Pro and ArcGIS Desktop installed, and it is important that you pick the right one from ArcGIS Pro using Python 3.
In the event that there isn’t a shortcut, you can start Python from a standard Windows command prompt by typing :
"%PROGRAMFILES%\ArcGIS\Pro\bin\Python\Scripts\propy"
The instructions above mirror Esri's help [6]for running Python.
Open your Python command prompt and you should be in the folder C:\Users\<username>\AppData\Local\ESRI\conda\envs\arcgispro-py3-clone\ or C:\Program Files\ArcGIS\Pro\bin\Python\envs\arcgispro-py3\ depending on your version of ArcGIS Pro. This is the default folder when the command prompt opens (which you can see at the prompt). Then type (and hit Enter) :
Scripts\pip install pyprof2calltree
Pip will then download from a repository of hosted Python packages the converter that we need and then install it. You should see a message saying “Successfully installed pyprof2calltree-1.4.3“ although your version number may be higher and that’s ok. If you receive an error message about permissions during the pyprof2calltree installation, close out of Python command prompt and reopen the program with administrative privileges (usually right-clicking on the program and selecting "Run as administrator," in Windows 10, you might need to "Open File Location" and then right-click to "Run as administrator").
After running commands in Python command prompt, you will probably also see an information message stating:
“You are using pip version 9.0.3, however version 10.0.1 is available. You should consider upgrading via the `python –m pip install –upgrade pip‘ command.“
You can ignore this message, we’re going to leave pip at its current version as that is what came with the ArcGIS Pro Python distribution and we know that it works.
1.7.2.3 Optional complementary materials: Code profiling with visualizations (continued)
This section is provided for interest only. Please feel free to skip over it and you can loop back to it at the end of the lesson if you have free time or after the end of the class.
Creating profile files
Now that we have the QCacheGrind viewer and converter installed we can create some profile files and we'll do that using IPython (available on another tab in Spyder). Think of IPython a little like the interactive Python window in ArcGIS with a few more bells and whistles. We’ll use IPython very shortly for line-by-line profiling, so as a bridge to that task, let's use it to perform some more simple profiling.
Open the IPython pane in Spyder and then you will probably need to change to the folder where your code is located. This should be as easy as looking at the top of the Spyder window, selecting and copying the folder name where your code is, then clicking in the IPython window and typing cd and pasting in the folder name and hitting enter (as seen below).
This might look like:
cd c:\Users\YourName\Documents\GEOG489\Lesson1\
We could (but won't) run our code from IPython by typing:
run Lesson1B_basic_raster_for_mp.py
and hitting enter and our code will run and display its output in the IPython window.
That is somewhat useful, but more useful is the ability to use additional packages within IPython for more detailed profiling. Let’s create a function-level profile file using IPython for that raster code and then we’ll run it through the converter and then visualize it in QCacheGrind.
To create that function-level profile we’ll use the built-in profiler prun.
We’ll access it using is a magic word instruction to Spyder which is shorthand for loading an external package; that package is called line_profiler which we just installed. Think of it in the same way as import for Python code – we’re now going to be able to access functionality embedded within that package.
Our magic words have % in front of them. Let's use it to see what the parameters are for using prun with (the ? on the end tells prun to show us its built-in help):
%prun?
If you scroll back up through the IPython console, you will be able to see all of the options for prun. You can compare our command below to that list to break down the various options and experiment with others if you wish. Notice the very last line of the help which states:
If you want to run complete programs under the profiler's control, use "%run -p [prof_opts] filename.py [args to program]" where prof_opts contains profiler specific options as described here.
That is what we’re going to do – use run with the prun options.
cd "C:\Users\YourName" %run -p -T profile_run.txt -D profile_run.prof Lesson1B_basic_raster_for_mp
There are a couple of important things to note in these commands. The first is the double quotes around the full path name in the cd command – these are important: just in case there is a space in your path, the double quotes encapsulate it so your folder is found correctly. The other thing is the casing of the various parameters (remember Python is case-sensitive and so are a lot of the built-in tools).
It could take a little while to complete our profiling as our code will run through from start to end. We can check that our code is running by opening the Windows Task Manager and watching the CPU usage which is probably at 100% on one of our cores.
While our code is running we’ll see the normal output with the timing print functions we had implemented earlier. When the run command completes we’ll see a few lines of output that look like :
%run -p -T profile_run.txt -D profile_run.prof
Lesson1B_basic_raster_for_mp
*** Profile stats marshalled to file 'profile_run.prof'.
*** Profile printout saved to text file 'profile_run.txt'.
3 function calls in 0.000 seconds
Ordered by: internal time
ncalls tottime percall cumtime percall
filename:lineno(function)
1 0.000 0.000 0.000 0.000 {built-in method
builtins.exec}
1 0.000 0.000 0.000 0.000
...
1 0.000 0.000 0.000 0.000
SSL.py:677(Session)
1 0.000 0.000 0.000 0.000
cookiejar.py:1753(LoadError)
1 0.000 0.000 0.000 0.000
socks.py:127(ProxyConnectionError)
1 0.000 0.000 0.000 0.000
_conditional.py:177(cryptography_has_mem_functions)
1 0.000 0.000 0.000 0.000
_conditional.py:196(cryptography_has_x509_store_ctx_get_issuer)
1 0.000 0.000 0.000 0.000
_conditional.py:210(cryptography_has_evp_pkey_get_set_tls_enco
dedpoint)
These summary outputs will also be written to a text file that we can open in a text editor (or in Spyder) and our profile file which we will convert and open in QCacheGrind. Writing that output to a text file is useful because there is too much of it to fit within IPython’s window buffer, and you won’t be able to get back to the output right at the start of the execution. If you open the profile_run.txt file in Spyder you’ll see the full output.
Convert output profile files with pyprof2calltree
We’ll run the converter using some familiar Python commands and the convert function within the IPython window:
from pyprof2calltree import convert, visualize
convert('profile_run.prof','callgrind.profile_run')
Open the converted files in QCacheGrind and inspect graphs
The converted output file can now be opened in QCacheGrind. Open QCacheGrind from the folder you installed it into earlier by double-clicking its icon. Click the folder icon in the top left of QCacheGrind or choose File ->Open from the menu and open the callgrind.profile_run file we just created, which should be in the same folder as your source code.
What we have now is a complicated and detailed interface and visualization of every function that our code called but in a more graphically friendly format than the original text file. We can sort by time, number of times a function was called, the function name and the location of that code (our own, within the arcpy library or another library) in the left-hand pane of the interface.
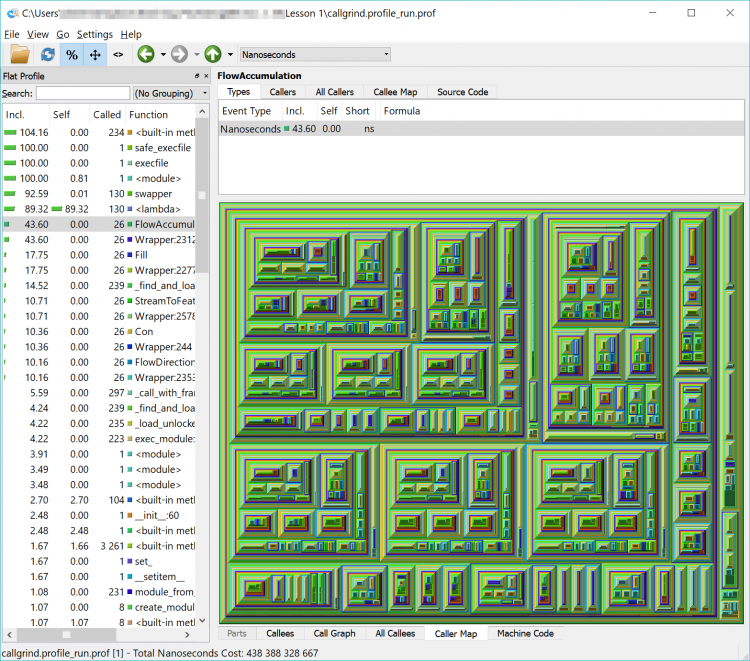
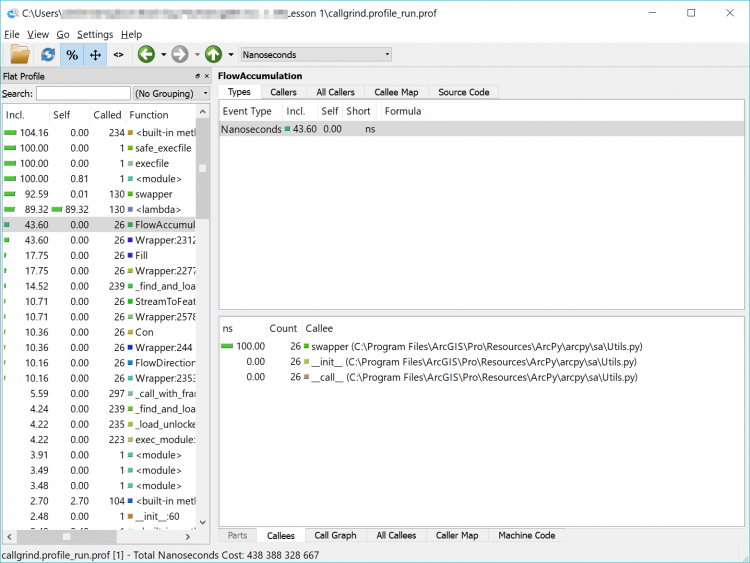
In the list, you will see a lot of built-in functions (things that Python does behind the scenes or that arcpy has it do – calculations, string functions etc.) but you will also see the names of some of the arcpy functions that we used such as FlowAccumulation(...) or StreamToFeature(...). If you double-click on one of them and click on the Call Graph tab in the lower pane you will see a graphical representation of where the tasks were called from. If you double-click on the function’s box above it in the Call Graph pane
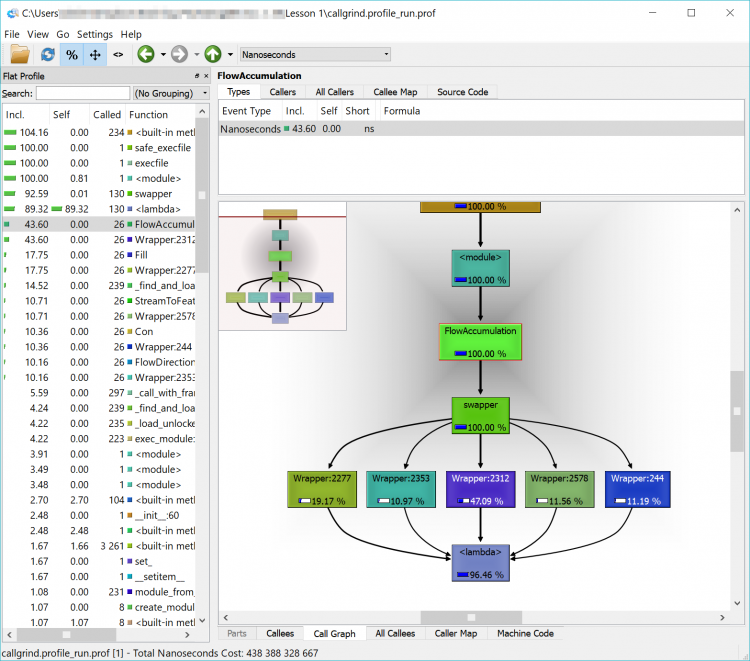
In the example below, we can see that FlowAccumulation(...) is the slowest of our tasks taking about 43% of the execution time. If we can find a way to speed up (or eliminate this process), we’ll make our code more efficient. If we can’t – that’s ok too – we’ll just have to accept that our code takes a certain amount of time to run.
Spend a little time clicking around in the interface and exploring your code – don’t worry too much about going down the rabbit hole of optimizing your code – just explore. Check out functions whose name you recognize such as those raster ones, or ListRasters(). Experiment with examining the content of the different tabs and seeing which modules call which functions (Callers, All Callers and Callee Map). Click down deep into one of the modules and watch the Callee Map change to show each small task being undertaken. If you get too far down the rabbit hole you can use the up arrow near the menu bar to find your way back to the top level modules.
If you’re interested, feel free to run through the process again with your multiprocessing code and see the differences. As a quick reminder, the IPython commands are (although your filenames might be different and be sure to double-check that you're in the correct folder if you get file not found errors):
%run -p -T profile_run_mp.txt -D profile_run_mp.prof Lesson1B_basic_raster_using_mp
from pyprof2calltree import convert, visualize
convert('profile_run_mp.prof','callgrind.profile_run_mp')
If you load that newly created file into QCacheGrind, you’ll note it looks a little different – like an Escher drawing or a geometric pattern. That is the representation of the multiprocessing functions being run. Feel free to explore among here as well – and you will notice that the functions we previously saw are harder to find – or invisible.
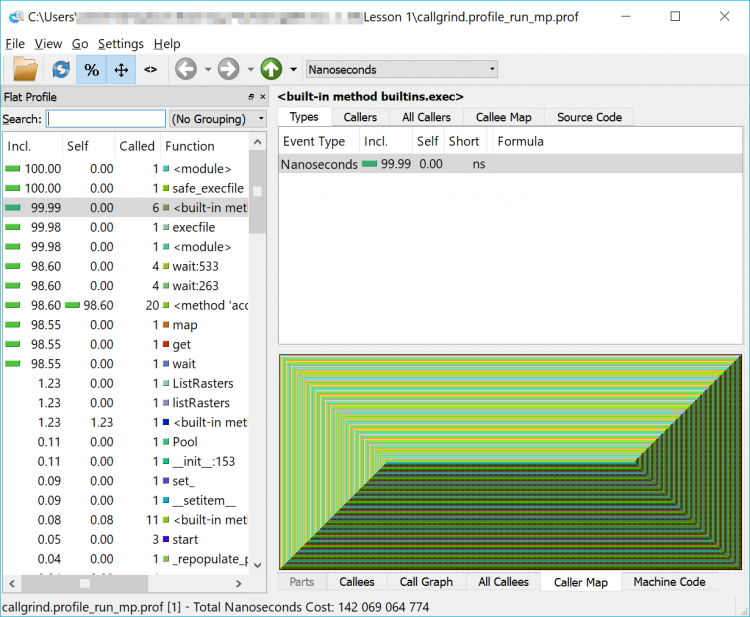
I haven't forgotten about my promise from earlier in the lesson to review the reasons why the Cherry-O code is only about 2-3x faster in multiprocessing mode than the 4x that we would have hoped.
Feel free to run both versions of your Cherry-O code against the profiler and you'll notice that most of the time is taken up by some code described as {method 'acquire' of '_thread.lock' objects} which is called a small number of times. This doesn't give us a lot of information but does hint that perhaps the slower performance is related to something to do with handling multiprocessing objects.
Remember back to our brief discussion about pickling objects which was required for multiprocessing?
It's the culprit, and the following optional section on line profiling will take a closer look at this issue. However, as we said before, line profiling adds quite a bit of complexity, so feel free to skip this section entirely or get back to it after you have worked through the rest of the lesson.
1.7.2.4 Optional complementary materials: Line profiling
As we said at the beginning of this section, this section is provided for interest only. Please feel free to skip over it and you can loop back to it at the end of the lesson if you have free time or after the end of the class. Be warned: installing the required extensions and using them is a little complicated - but this is an advanced class, so don't let this deter you.
Before we begin this optional section, you should know that line profiling is slow – it adds a lot of overhead to our code execution and it should only be done on functions that we know are slow and we’re trying to identify the specifics of why they are slow. Due to that overhead, we cannot rely on the absolute timing reported in line profiling, but we can rely on the relative timings. If a line of code is taking 50% of our execution time and we can reduce that to 40,30 or 20% (or better) of our total execution time then we have been successful.
With that warning about the performance overhead of line profiling, we’re going to install our line profiler (which isn’t the same one that Spyder or ArcGIS Pro would have us install) again using pip (although you are welcome to experiment with those too).
Setting PermissionsBefore we start installing packages we will need to adjust operating system (Windows) permissions on the Python folder of ArcGIS Pro, as we will need the ability to write to some of the folders it contains. This will also help us if we inadvertently attempt to create profile files in the Python folder as the files will be created instead of producing an error that the folder is inaccessible (but we shouldn't create those files there as it will add clutter).
Open up Windows Explorer and navigate to the c:\Program Files\ArcGIS\Pro\bin folder. Select the Python folder and right click on it and select Properties. Select the Security tab, and click the Advanced button. In the new window that opens select Change Permissions, Select the Users group, Uncheck the Include inheritable permissions from this object’s parent box or Disable Inheritance – depending on your version of Windows, and select Add (or Make Explicit) on the dialog that appears.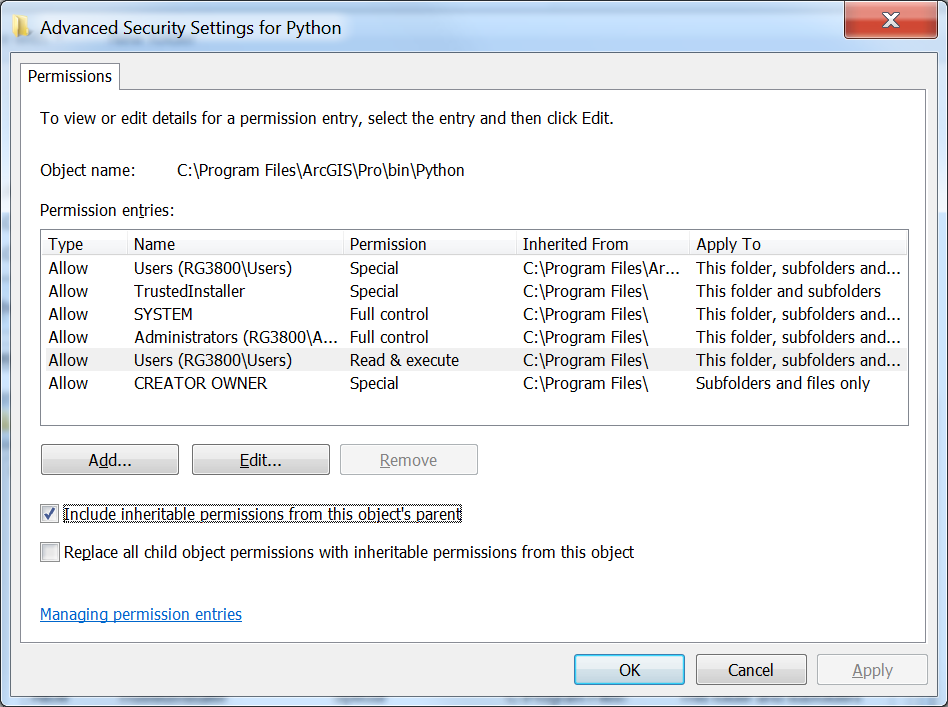
Once again open (if it isn’t already) your Python command prompt and you should be in the folder C:\Program Files\ArcGIS\Pro\bin\Python\envs\arcgispro-py3 or C:\Users\<username>\AppData\Local\ESRI\conda\envs\arcgispro-py3-clone, the default folder of your ArcGIS Pro Python environment, when that command prompt opens (which you can see at the prompt). If in your version of Pro, the command prompt instead shows C:\Users\<username>\AppData\Local\ESRI\conda\envs\arcgispro-py3-clone, then you will have to use this path instead in some of the following commands where the kernprof program is used.
Then type :
scripts\pip install line_profiler
Pip will then download from a repository of hosted Python packages the line_profiler that we need and then install it.
If you receive an error that "Microsoft Visual C++ 14.0 is required," visit https://www.visualstudio.com/downloads/ [7] and download the package for "Visual Studio Community 2017" which will download Visual Studio Installer. Run Visual Studio Installer, and under the "Workloads" tab, you will select two components to install. Under Windows, check the box in the upper right hand corner of "Desktop Development with C++," and, under Web & Cloud, check the box for "Python development." After checking the boxes, click "Install" in the lower right hand corner. After installing those, open the Python command prompt again and enter:
scripts\pip install misaka
If that works, then install the line_profiler with...
scripts\pip install line_profiler
You should see a message saying "Successfully installed line_profiler 2.1.2" although your version number may be higher and that’s okay.
Now that IPython is aware of the line profiler we can run it. There are two modes for running the line profiler, function mode, where we supply a Python file and a function we want to run as well as the parameters for that function given as parameters to the profiler, and module mode, where we supply the module name.
Function-level line profiling is very useful when you want to test just a single function, or if you’re doing multiprocessing as we saw above. Module-level line profiling is a useful first pass to identify those functions that might be slowing things down and it's why we did a similar approach with our higher-level profiling earlier.
Now we can dive into function-level profiling to find the specific lines which might be slowing down our code and then optimize or enhance them and then perform further function-level line profiling (or module-level profiling) again to test our improvements.
We will start with module-level profiling using our single processor Cherry-O code [8], look at our non-multiprocessing raster example code [9] that did the analysis of the Penn State campus, and finally move on to the multiprocessing Cherry-O code [10]. You may notice a few little deviations in the code that is being used in this section compared to the versions presented in Section 1.5 and 1.6. These deviations are really minimal and have no effect on how the code works and the insights we gain from the profiling.
Our line profiler is in a package called KernProf (named after its author) and it works as a wrapper around the standard cProfile and Line_Profiler tools.
We need to make some changes to our code so that the line profiler knows which functions we wish it to interrogate. The first of those changes is to wrap a function definition around our code (so that we have a function to profile instead of just a single block of code). The second change we need is to use a decorator which is an instruction or meta information for a piece of code that will be ignored by Python. In the case of our profiler, we need to use the decorator @profile to tell KernProf which functions (and we can do many) to examine and it will ignore any without a decorator. Your decorator may give you errors if you're not running your code against the profiler so in that case comment it out.
We’ve made those changes to our original Cherry-O code below so you can see them for yourselves. Check out line 8 for the decorator, line 9 for the function definition (and note how the code is now indented within the function) and line 53 where the function is called. You might also notice that I reduced the number of games back down to 10001 from our very large number earlier. Don’t forget to save your code after you make these changes.
# Simulates 10K game of Hi Ho! Cherry-O
# Setup _very_ simple timing.
import time
start_time = time.time()
import random
@profile
def cherryo():
spinnerChoices = [-1, -2, -3, -4, 2, 2, 10]
turns = 0
totalTurns = 0
cherriesOnTree = 10
games = 0
while games < 10001:
# Take a turn as long as you have more than 0 cherries
cherriesOnTree = 10
turns = 0
while cherriesOnTree > 0:
# Spin the spinner
spinIndex = random.randrange(0, 7)
spinResult = spinnerChoices[spinIndex]
# Print the spin result
#print ("You spun " + str(spinResult) + ".")
# Add or remove cherries based on the result
cherriesOnTree += spinResult
# Make sure the number of cherries is between 0 and 10
if cherriesOnTree > 10:
cherriesOnTree = 10
elif cherriesOnTree < 0:
cherriesOnTree = 0
# Print the number of cherries on the tree
#print ("You have " + str(cherriesOnTree) + " cherries on your tree.")
turns += 1
# Print the number of turns it took to win the game
#print ("It took you " + str(turns) + " turns to win the game.")
games += 1
totalTurns += turns
print ("totalTurns "+str(float(totalTurns)/games))
#lastline = raw_input(">")
# Output how long the process took.
print ("--- %s seconds ---" % (time.time() - start_time))
cherryo()
We could try to run the profiler and the other code from within IPython but that often causes issues such as unfound paths, files, etc., as well as making it difficult to convert our output to a nice readable text file. Instead, we’ll use our Python command prompt and then we’ll run the line profiler using (note: the "-l" is a lowercase L and not the number 1):
python "c:\program files\arcgis\pro\bin\python\envs\arcgispro-py3\lib\site-packages\kernprof.py" -l “c:\users\YourName\CherryO.py”
When the profiling completes (and it will be fast in this simple example), you’ll see our normal code output from our print functions and the summary from the line profiler:
Wrote profile results to CherryO.py.lprof
This tells us the profiler has created an lprof file in our current directory called CherryO.py.lprof (or whatever our input code was called).
The profile files will be saved wherever your Python command prompt path is pointing. Unless you've changed the directory, the Python command prompt will most likely be pointing to C:\Program Files\ArcGIS\Pro\Bin\Python\envs\arcgispro-py3, and the files will be saved in that folder.
The profile files are binary files that will be impossible for us to read without some help from another tool. So to rectify that we’ll run that .lprof file through the line_profiler (which seems a little confusing because you would think we just created that file with the line_profiler and we did, but the line_profiler can also read the files it created) and then pipe (redirect) the output to a text file which we’ll put back in our code directory so we can find it more easily.
To achieve this, we run the following command in our Python command window:
..\python –m line_profiler CherryO.py.lprof > "c:\users\YourName\CherryO.profile.txt"
This command will instruct Python to run the line_profiler (which is some Python code itself) to process the .lprof file we created. The > will redirect the output to a text file at the provided path instead of displaying the output to the screen.
We can then open the resulting output file which should be back in our code folder from within Spyder and read the results. I’ve included my output for reference below and they are also in the CherryO.Profile pdf [11].
Timer unit: 4.27655e-07 s
Total time: 3.02697 s
File: c:\users\YourName\Lesson 1\CherryO.py
Function: cherryo at line 8
Line # Hits Time Per Hit % Time Line Contents
==============================================================
8 @profile
9 def cherryo():
10 1 5.0 5.0 0.0 spinnerChoices = [-1, -2, -3, -4, 2, 2, 10]
11 1 2.0 2.0 0.0 turns = 0
12 1 1.0 1.0 0.0 totalTurns = 0
13 1 1.0 1.0 0.0 cherriesOnTree = 10
14 1 1.0 1.0 0.0 games = 0
15
16 10002 36775.0 3.7 0.5 while games < 10001:
17 # Take a turn as long as you have more than 0 cherries
18 10001 25568.0 2.6 0.4 cherriesOnTree = 10
19 10001 27091.0 2.7 0.4 turns = 0
20 168060 464529.0 2.8 6.6 while cherriesOnTree > 0:
21
22 # Spin the spinner
23 158059 4153276.0 26.3 58.7 spinIndex = random.randrange(0, 7)
24 158059 487698.0 3.1 6.9 spinResult = spinnerChoices[spinIndex]
25
26 # Print the spin result
27 #print "You spun " + str(spinResult) + "."
28
29 # Add or remove cherries based on the result
30 158059 460642.0 2.9 6.5 cherriesOnTree += spinResult
31
32 # Make sure the number of cherries is between 0 and 10
33 158059 458508.0 2.9 6.5 if cherriesOnTree > 10:
34 42049 112815.0 2.7 1.6 cherriesOnTree = 10
35 116010 325651.0 2.8 4.6 elif cherriesOnTree < 0:
36 5566 14506.0 2.6 0.2 cherriesOnTree = 0
37
38 # Print the number of cherries on the tree
39 #print "You have " + str(cherriesOnTree) + " cherries on your tree."
40
41 158059 445969.0 2.8 6.3 turns += 1
42 # Print the number of turns it took to win the game
43 #print "It took you " + str(turns) + " turns to win the game."
44 10001 29417.0 2.9 0.4 games += 1
45 10001 31447.0 3.1 0.4 totalTurns += turns
46
47 1 443.0 443.0 0.0 print ("totalTurns "+str(float(totalTurns)/games))
48 #lastline = raw_input(">")
49
50 # Output how long the process took.
51 1 3723.0 3723.0 0.1 print ("--- %s seconds ---" % (time.time() - start_time))
What we can see here are the individual times to run each line of code: The numbers on the left are the code line numbers, the number of times each line was run (Hits), the time each line took in total (Hits * Time Per Hit), the time per hit, the percentage of time those lines took and, for reference, the line of code alongside.
The first thing that jumps out at me is that the random number selection (line 23) takes the longest time and is called the most – if we can speed this up somehow we can improve our performance.
Let’s move onto the other examples of our code to see some more profiling results.
First we’ll look at the sequential version of our raster processing code before coming back to look at the multiprocessing examples as they’re a little special.
As before with the Cherry-O example we’ll need to wrap our code into a function and use the @profile decorator (and of course call the function). Attempt to make these changes on your own and if you get stuck you can find my code sample here [12] if you want to check your work against mine.
We’ll run the profiler again, produce our output file and then convert it to text and review the results using:
python "c:\program files\arcgis\pro\bin\python\envs\arcgispro-py3\lib\site-packages\kernprof.py" -l "c:\users\YourName\Lesson1B_basic_raster_for_mp.py"
and then
..\python –m line_profiler Lesson1B_basic_raster_for_mp.py.lprof > "c:\users\YourName\Lesson1B_basic_raster_for_mp.py_profile.txt"
If we investigate these outputs (my outputs are here [13]) we can see that the Flow Accumulation calculation is again the slowest, just as we saw when we were doing the module-level calculations. In this case, because we’re predominantly using arcpy functions, we’re not seeing as much granularity or resolution in the results. That is, we don’t know why Flow Accumulation(...) is so slow but I’m sure you can see that in some other circumstances, you could identify multiple arcpy functions which could achieve the same result – and choose the most efficient.
Next, we’ll look at the multiprocessing example of the Cherry-O example to see how we can implement line profiling into our multiprocessing code. As we noted earlier, multiprocessing and profiling are a little special as there is a lot going on behind the scenes and we need to very carefully select what functions and lines we’re profiling as some things cannot be pickled.
Therefore what we need to do is use the line profiler in its API mode. That means instead of using the line profiling outside of our code, we need to embed it in our code and put it in a function between our map and the function we’re calling. This will give us output for each process that we launch. Now if we do this for the Cherry-O example we’re going to get 10,000 files – but thankfully they are small so we’ll work through that as an example.
The point to reiterate before we do that is the Cherry-O code runs in seconds (at most) – once we make these line profiling changes the code will take a few minutes to run.
We’ll start with the easy steps and work our way up to the more complicated ones:
import line_profiler
Now let’s define a function within our code to sit between the map function and the called function (cherryO in my version).
We’ll break down what is happening in this function shortly and that will also help to explain how it fits into our workflow. This new function will be called from our mp_handler() function instead of our original call to cherryO and this new function will then call cherryO.
Our new mp_handler function looks like:
def mp_handler():
myPool = multiprocessing.Pool(multiprocessing.cpu_count())
## The Map function part of the MapReduce is on the right of the = and the Reduce part on the left where we are aggregating the results to a list.
turns = myPool.map(worker,range(numGames))
# Uncomment this line to print out the list of total turns (but note this will slow down your code's execution
#print(turns)
# Use the statistics library function mean() to calculate the mean of turns
print(mean(turns))
Note that our call to myPool.map now has worker(...) not cherryO(...) as the function being spawned multiple times. Now let's look at this intermediate function that will contain the line profiler as well as our call to cherryO(...).
def worker(game):
profiler=line_profiler.LineProfiler(cherryO)
call = 'cherryO('+str(game)+')'
turns = profiler.run(call)
profiler.dump_stats('profile_'+str(game)+'.lprof')
return(turns)
The first line of our new function is setting up the line profiler and instructing it to track the cherryO(...) function.
As before, we pass the variable game to the worker(...) function and then pass it through to cherryO(...) so it can still perform as before. It’s also important that, when we call cherryO(...), we record the value it returns into a variable turns – so we can return that to the calling function so our calculations work as before. You will notice we’re not just calling cherryO(...) and passing it the variable though – we need to pass the variable a little differently as the profiler can only support certain picklable objects. The most straightforward way to achieve that is to encode our function call into a string (call) and then have the profiler run that call. If we don’t do this the profiler will run but no results will be returned.
Just before we send that value back we use the profiler’s function dump_stats to write out the profile results for the single game to an output file.
Don’t forget to save your code after you make these changes. Now we can run through a slightly different (but still familiar) process to profile and export our results, just with different file names. To run this code we’ll use the Python command prompt:
python c:\users\YourName\CherryO_mp.py
Notice how much longer the code now takes to run – this is another reason to wrap the line profiling in its own function. That means that we don’t need to leave it in production code; we can just change the function calls back and leave the line profiling code in place in case we want to test it again.
It's also possible you'll receive several error messages when the code runs, but the lprof files are still created.
Once our code completes, you will notice we have those 10,000 lprof files (which is overkill as they are probably all largely the same). Examine a few of the files if you like by converting them to text files and viewing them in your favorite text editor or Spyder using the following in the Python command prompt:
python –m line_profiler profile_1.lprof > c:\users\YourName\profile_1.txt
If you examine one of those files, you’ll see results similar to:
Timer unit: 4.27655e-07 s
Total time: 0.00028995 s
File: c:\users\obrien\Dropbox\Teaching_PSU\Geog489_SU_1_18\Lesson 1\CherryO_MP.py
Function: cherryO at line 25
Line # Hits Time Per Hit % Time Line Contents
==============================================================
25 def cherryO(game):
26 1 11.0 11.0 1.6 spinnerChoices = [-1, -2, -3, -4, 2, 2, 10]
27 1 9.0 9.0 1.3 turns = 0
28 1 8.0 8.0 1.2 totalTurns = 0
29 1 8.0 8.0 1.2 cherriesOnTree = 10
30 1 9.0 9.0 1.3 games = 0
31
32 # Take a turn as long as you have more than 0 cherries
33 1 9.0 9.0 1.3 cherriesOnTree = 10
34 1 9.0 9.0 1.3 turns = 0
35 16 41.0 2.6 6.0 while cherriesOnTree > 0:
36
37 # Spin the spinner
38 15 402.0 26.8 59.3 spinIndex = random.randrange(0, 7)
39 15 38.0 2.5 5.6 spinResult = spinnerChoices[spinIndex]
40
41 # Print the spin result
42 #print "You spun " + str(spinResult) + "."
43
44 # Add or remove cherries based on the result
45 15 34.0 2.3 5.0 cherriesOnTree += spinResult
46
47 # Make sure the number of cherries is between 0 and 10
48 15 35.0 2.3 5.2 if cherriesOnTree > 10:
49 4 8.0 2.0 1.2 cherriesOnTree = 10
50 11 24.0 2.2 3.5 elif cherriesOnTree < 0:
51 cherriesOnTree = 0
52
53 # Print the number of cherries on the tree
54 #print "You have " + str(cherriesOnTree) + " cherries on your tree."
55
56 15 32.0 2.1 4.7 turns += 1
57 # Print the number of turns it took to win the game
58 1 1.0 1.0 0.1 return(turns)
Arguably we’re not learning anything that we didn’t know from the sequential version of the code – we can still see the randrange() function is the slowest or most time consuming (by percentage) – however, if we didn’t have the sequential version and wanted to profile our multiprocessing code this would be a very important skill.
The same steps to modify our code above would be implemented if we were performing this line profiling on arcpy (or any other) multiprocessing code. The same type of intermediate function would be required, we would need to pass and return parameters (if necessary) and also reformat the function call so that it was picklable. The output from the line profiler is delivered in a different format to the module-level profiling we were doing before and, therefore, isn’t suitable for loading into QCacheGrind. I'd suggest that isn't as important as we're looking at a much smaller number of lines of code, so the graphical representation isn't as important.
Returning to our ongoing discussion about the less than anticipated performance improvement between our sequential and multiprocessing Cherry-O code, what we can infer by comparing the line profile output of the sequential version of our code and the multiprocessing version is that pretty much all of the steps take the same proportion of time. So if we're doing nearly everything in about the same proportions, but 4 times as many of them (using our 4 processor PC example) then why isn't the performance improvement around 4x faster? We'd expect that setting up the multiprocessing environment might be a little bit of an overhead so maybe we'd be happy with 3.8x or so.
That isn't the case though so I did a little bit of experimenting with calculating how much time it takes to pickle those simple integers. I modified the mp_handler function in my multiprocessor code so that instead of doing actual work selecting cherries, it pickled the 1 million integers that would represent the game number. That function looked like this (nothing else changed in the code):
import pickle
def mp_handler():
myPool = multiprocessing.Pool(multiprocessing.cpu_count())
## The Map function part of the MapReduce is on the right of the = and the Reduce part on the left where we are aggregating the results to a list.
#turns = myPool.map(worker,range(numGames))
#turns = myPool.map(cherryO,range(numGames))
t_start=time.time()
for i in range(0,numGames):
pickle_data = pickle.dumps(range(numGames))
print ("cPickle data took",time.time()-t_start)
t_start=time.time()
pickle.loads(pickle_data)
print ("cPickle data took",time.time()-t_start)
What I learned from this experimentation was that the pickling took around 4 seconds - or about 1/4 of the time my code took to play 1M Cherry-O games in multiprocessing mode - 16 seconds (it was 47 seconds for the sequential version).
A simplistic analysis suggests that the pickling comprises about 25% of my execution time (and your results might vary). If I subtract the time taken for the pickling, my code would have run in 12 seconds and then 47s ÷ 12s = 3.916 - or the nearly the 4x improvement we would have anticipated. So the takeaway message here is the reinforcing of some of the implications of implementing multiprocessing that we discussed earlier: there is an overhead to multiprocessing and lots of small calculations as in this case aren't the best application for it because we lose some of that performance benefit due to that implementation overhead. Still, an almost tripling of performance (47s / 16s) is worth the effort.
Your coding assessment for this lesson will have you modify some code (as mentioned earlier) and profile some multiprocessing analysis code which is why we’re not demonstrating it specifically here. See the lesson assignment page for full details.
1.7.2.5 A last word on profiling
Before we move on from profiling a few important points need to be made. As you might have worked out for yourself by this point, profiling is time-consuming and you really should only undertake it if you have a very slow piece of code or one where you will be running it thousands of times or more and a small performance improvement is likely to be beneficial in the long run. To put that another way, if you spend a day profiling your code to improve its performance and you reduce the execution time from ten minutes to five minutes but you only run that code once then I would argue you haven’t used your time productively. If your code is already fast enough that it executes in a reasonable amount of time – that is fine.
Do not get caught in the trap of beginning to optimize and profile your code too early, particularly before the code is complete. You may be focusing on a slow piece of code that will only be executed once or twice and the performance improvement will not be significant compared to the execution of the other 99% of the code.
We have to accept that some external libraries are inefficient and if we need to use them, then we must accept that they take as long as they do to get the job done. It is also possible that those libraries are extremely efficient and take as long as they do because the task they are performing is complicated. There isn’t any point attempting to speed up the arcpy.da cursors for example as they are probably as fast as they are likely to be in the near future. If that is the slowest part of our code, we may have to accept that.